September 15 2017
In RNA-seq gene expression data analysis, we come across various expression units such as RPM, RPKM, FPKM and raw reads counts. Most of the times it's difficult to understand basic underlying methodology to calculate these units from mapped sequence data. Here I attempted to explain these units in much simpler way.
Why different normalized expression units:
The expression units provide a digital measure of the abundance of transcripts. Normalized expression units are necessary to remove technical biases in sequenced data such as depth of sequencing (more sequencing depth produces more read count for gene expressed at same level) and gene length (differences in gene length generate unequal reads count for genes expressed at the same level; longer the gene more the read count).
Gene expression units and calculation:
RPM (Reads per million mapped reads)
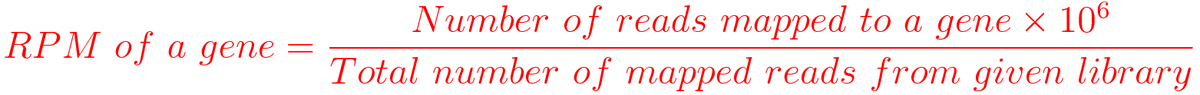
For example, You have sequenced one library with 5 million(M) reads. Among them, total 4 M matched to the genome sequence and 5000 reads matched to a given gene.
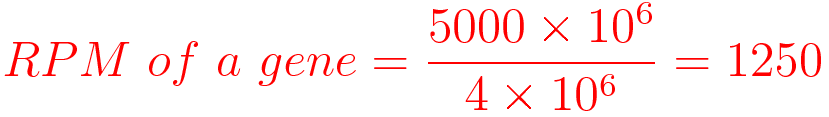
Notes:
RPKM (Reads per kilo base per million mapped reads)

Here, 103 normalizes for gene length and 106 for sequencing depth factor.
FPKM (Fragments per kilo base per million mapped reads) is analogous to RPKM and used especially in paired-end RNA-seq experiments. In paired-end RNA-seq experiments, two (left and right) reads are sequenced from same DNA fragment. When we map paired-end data, both reads or only one read with high quality from a fragment can map to reference sequence. To avoid confusion or multiple counting, the fragments to which both or single read mapped is counted and represented for FPKM calculation.
For example, You have sequenced one library with 5 M reads. Among them, total 4 M matched to the genome sequence and 5000 reads matched to a given gene with a length of 2000 bp.

Notes:
TPM (Transcript per million)
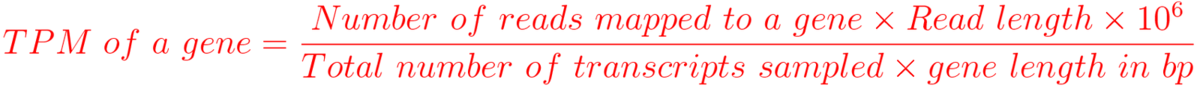
Here, read length refers to the average number of nucleotides mapped to a gene.
For example, You have sequenced one library with 5M 100 bp reads. Among them, total 4M matched to the genome sequence and 5000 reads matched to a given gene with a length of 2000 bp. There were 10K transcripts were sampled from a genome sequence i.e. reads mapped to 10K genes. Suppose all 100 bp mapped from 5000 reads.

Notes:
Relationship between RPKM and TPM,

References: